Simulation of a compartmental infectious disease transmission model illustrating different types of direct transmission
Source:R/simulate_directtransmission_ode.R
simulate_directtransmission_ode.RdThis model allows for the simulation of different direct transmission modes
simulate_directtransmission_ode(
S = 999,
I = 1,
bd = 0.005,
bf = 0,
A = 2,
n = 0,
m = 0,
g = 0.1,
w = 0,
scenario = 1,
tmax = 120
)Arguments
- S
: initial number of susceptibles : numeric
- I
: initial number of infected hosts : numeric
- bd
: rate of transmission for density-dependent transmission : numeric
- bf
: rate of transmission for frequency-dependent transmission : numeric
- A
: the size of the area in which the hosts are assumed to reside/interact : numeric
- n
: the rate of births : numeric
- m
: the rate of natural deaths : numeric
- g
: the rate at which infected hosts recover : numeric
- w
: the rate of waning immunity : numeric
- scenario
: choice between density dependent (=1) and frequency dependent (=2) transmission : numeric
- tmax
: maximum simulation time, units of months : numeric
Value
This function returns the simulation result as obtained from a call to the deSolve ode solver.
Details
A compartmental ID model with several states/compartments is simulated as a set of ordinary differential equations. The function returns the output from the odesolver as a list, with the element ts, which is a dataframe whose columns represent time, the number of susceptibles, the number of infected, and the number of recovered.
Warning
This function does not perform any error checking. So if you try to do something nonsensical (e.g. any negative values or fractions > 1), the code will likely abort with an error message
References
See e.g. Keeling and Rohani 2008 for SIR models and the documentation for the deSolve package for details on ODE solvers
See also
The UI of the Shiny app 'DirectTransmission', which is part of this package, contains more details on the model.
Examples
# To run the simulation with default parameters just call this function:
result <- simulate_directtransmission_ode()
# To choose parameter values other than the standard one, specify them like such:
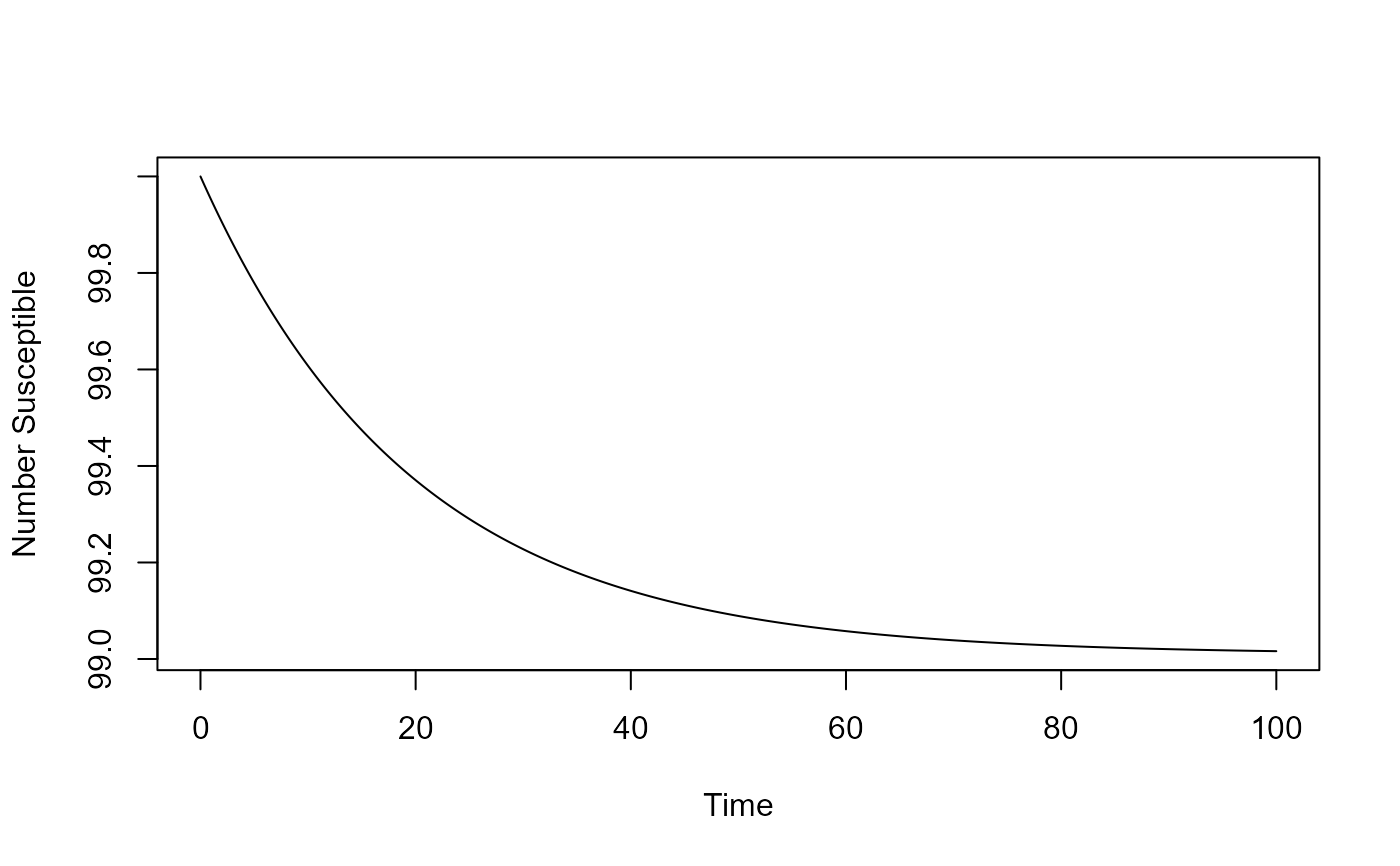
result <- simulate_directtransmission_ode(S = 100, tmax = 100, A=10)
# You should then use the simulation result returned from the function, like this:
plot(result$ts[,"time"],result$ts[,"S"],xlab='Time',ylab='Number Susceptible',type='l')